ChatGPT for Medicine: Transforming Healthcare With LLMs
A research team supported by Nvidia built a new large language model specifically for healthcare. They basically created a ChatGPT-equivalent for the clinical setting.

Although large language models (LLMs) were around before, their popularity grew since the introduction of ChatGPT in late 2022. Around that same time, a research group set out to develop a new LLM specific to healthcare. It's published in npj Digital Medicine and has since received almost 200 citations. Nvidia helped build the model and called it GatorTron. To grasp the motivation behind it, let's first understand its impact on healthcare.
Electronic Health Records
Electronic health records (EHR) are a more efficient way of storing patient data. At the same time, they present an opportunity to develop programs that analyse their data. The end-goal is to then use them in a clinical setting.
Generally, EHR contain structured and unstructured data. Structured data are much easier to analyse. They contain information such as patient demographics, disease codes, lab results and visit dates. Despite their usefulness, they often present an extra burden for medical staff. Generally, doctors lean more towards unstructured data, such as clinical narratives. And as you might imagine, they're much harder to analyse.
The relation between NLP and LLM
Fortunately, we have natural language processing (NLP) and LLMs. Using them, computers can understand human language, such as clinical narratives.
NLP is a way for computers to understand and interpret human language. Using NLP methods, computer scientists are then able to create (or, train) LLMs. They, in turn, are designed to work with human language at scale.
In a nutshell, LLMs are specific applications of the NLP methods and can be used in a variety of fields. For example, ChatGPT is also an LLM, but a very general one. Several biomedical LLMs are also available. Examples are BioBERT, BioMegatron and Clinical BERT, which were all included in the paper.
Training GatorTron
The aim of the paper was to develop a new clinical LLM. Additionally, the researchers tested to see how the number of parameters affects its performance and how it compares to existing models.
It's worth mentioning that NLP and LLMs are based on transformer architecture. It's a fancy way of saying the model can focus on different parts of the input when producing an output. Creating an LLM consists of pretraining and fine-tuning.
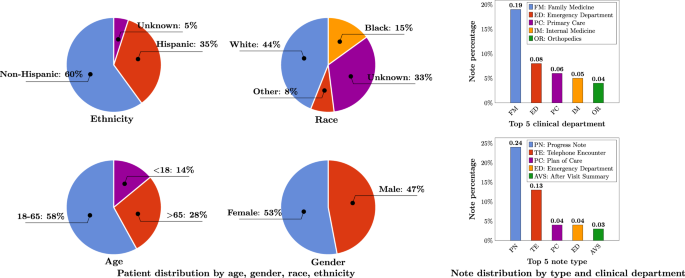
The research team used the University of Florida Health records to extract almost 300 million clinical notes from 2.4+ million patients. After preprocessing and de-identifying the data, they were left with about 82 billion medical words. They also added some from MIMIC-III (a critical care database), PubMed and Wikipedia. The final count was >90 billion words.
In comparison, Clinical BERT was trained using "only" 0.5 billion words. But the true test of GatorTron's capabilities lies in testing its clinical performance.
These 90 billion words were used to pretrain GatorTron and then fine-tune it for 5 different clinical NLP tasks:
- Scale up the size of training data and the number of parameters.
- Recognize clinical concepts and medical relations - identify concepts with important clinical meaning and classify them.
- Assess semantic textual similarity - determine the extent to which two sentences are similar.
- Natural language inference (NLI) - determine if a conclusion can be inferred from a given sentence.
- Medical question answering (MQA) - understand information from the entire clinical document.
They used 3 different settings with different number of parameters:
- Base model: 345 million
- Medium model: 3.9 billion
- Large model: 8.9 billion.
The idea was that by scaling the number of parameters, its performance on complex tasks would also improve.
How did it do?
Scaling GatorTron from the base model to the large one brought about remarkable performance improvements. This was best seen in complex tasks such as NLI and MQA. It's interesting that the largest model performed best in all tasks except for one. The medium model was better only at assessing semantic textual similarities.
Perhaps unsurprisingly, it was also better at all tasks than the alternatives. But in complex tasks such as NLI and MQA, even the largest model had difficulties in identifying key pieces of information. Overall, developing specific LLMs for clinical setting can yield useful applications for physicians. The whole LLM is also publicly available.
It would be interesting to test it against ChatGPT. It is more general, but also trained using a lot more words and parameters. An idea for a follow-up study?
This summary is based on research from the article A large language model for electronic health records by Yang, X., Chen, A., PourNejatian, N. et al. published in npj Digital Medicine. The original work is freely available and licensed under the Creative Commons Attribution 4.0 International Licence. Adaptations and summaries are provided by Medical Notes and are not endorsed by the original authors or publisher.





